On the occasion of the release of the LinkR article, we're taking a look back at LinkR’s progress one year after its acceleration with Data For Good.
LinkR and Data For Good – One year later
Introduction
In February 2024, the twelfth season of acceleration by Data For Good began. For three months, dozens of volunteers joined us to support the creation of an open-source, low-code, collaborative data science platform for healthcare.
Those three months were rich in discussions and collaboration, giving LinkR a huge boost. This led to a major redesign of the platform, making it more robust, more intuitive, and enriched with new plugins.
One year later, this collective work has materialized in a beautiful way: the publication of our article in the International Journal of Medical Informatics.
We’d like to take this opportunity to sincerely thank the members of Data For Good and the contributors of Season 12, and reflect on LinkR’s progress over the past year.
Website creation
Shortly after the end of Season 12, with the support of volunteers, we created the LinkR website.
This site hosts a comprehensive documentation designed both for clinicians with no programming skills and for data scientists or developers.
We created a Blog section, where we regularly publish content aimed at democratizing health data science—from explaining what a health data warehouse is to more technical guides like common OMOP SQL queries.
Lastly, an interactive demo lets you try the app online, with no installation required.
User interface redesign
InterHop is made up of clinicians, data scientists, and developers, enabling LinkR to evolve through close collaboration across disciplines.
This direct connection between end users and developers allowed us to rapidly design an intuitive interface tailored to real-world healthcare practices: exploring clinical data, building dashboards, creating analysis pipelines, etc.
Python integration
While R remains a reference language for statistical analysis, many essential data science libraries are now available only in Python.
That’s why we made it simple to write and run Python code directly within the LinkR interface. This integration significantly expands the platform’s capabilities, especially for creating custom plugins using tools like scikit-learn, PyTorch, or transformers.
Thanks to this compatibility, LinkR combines the best of both worlds: R and Python.
New plugin development
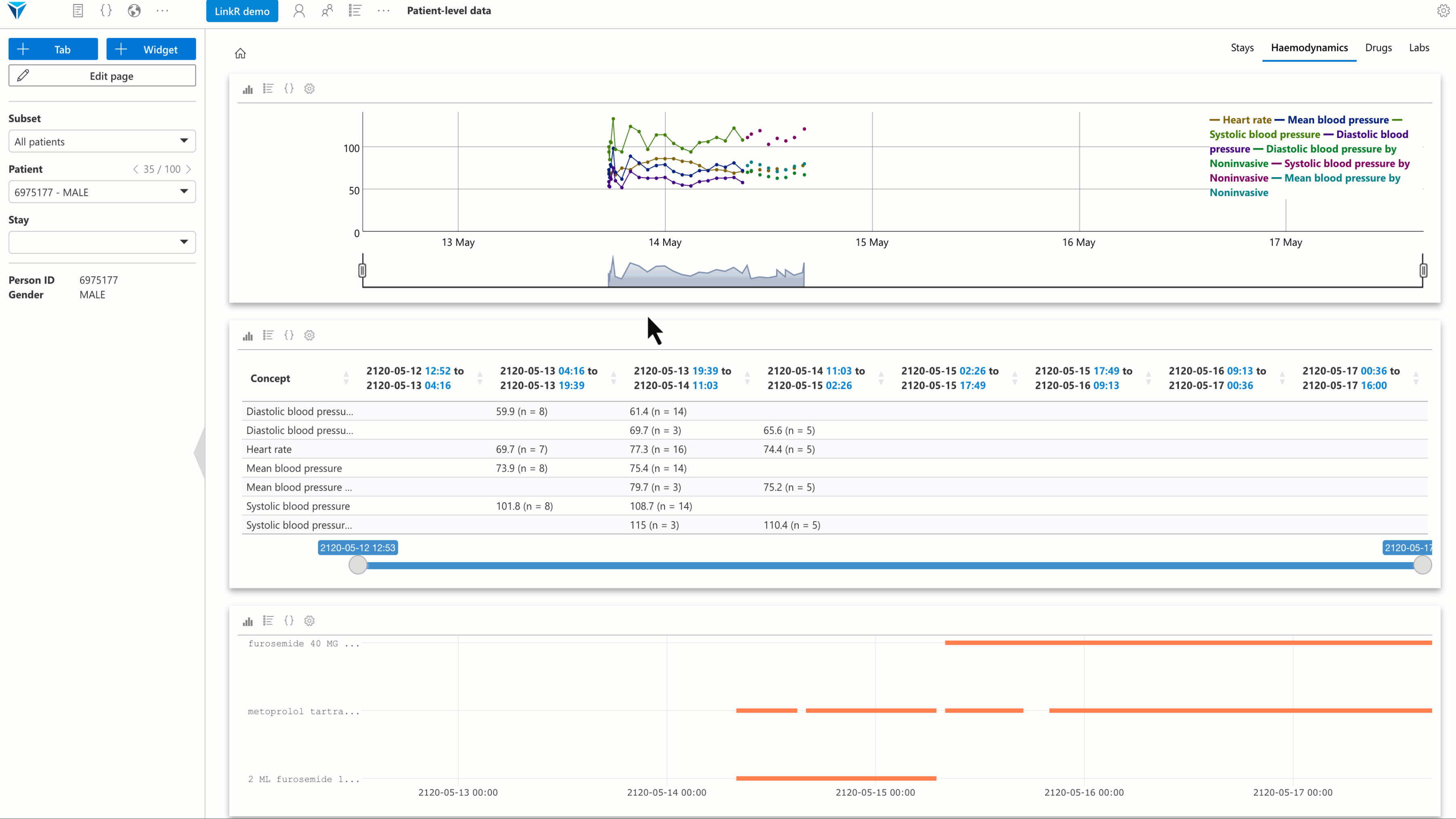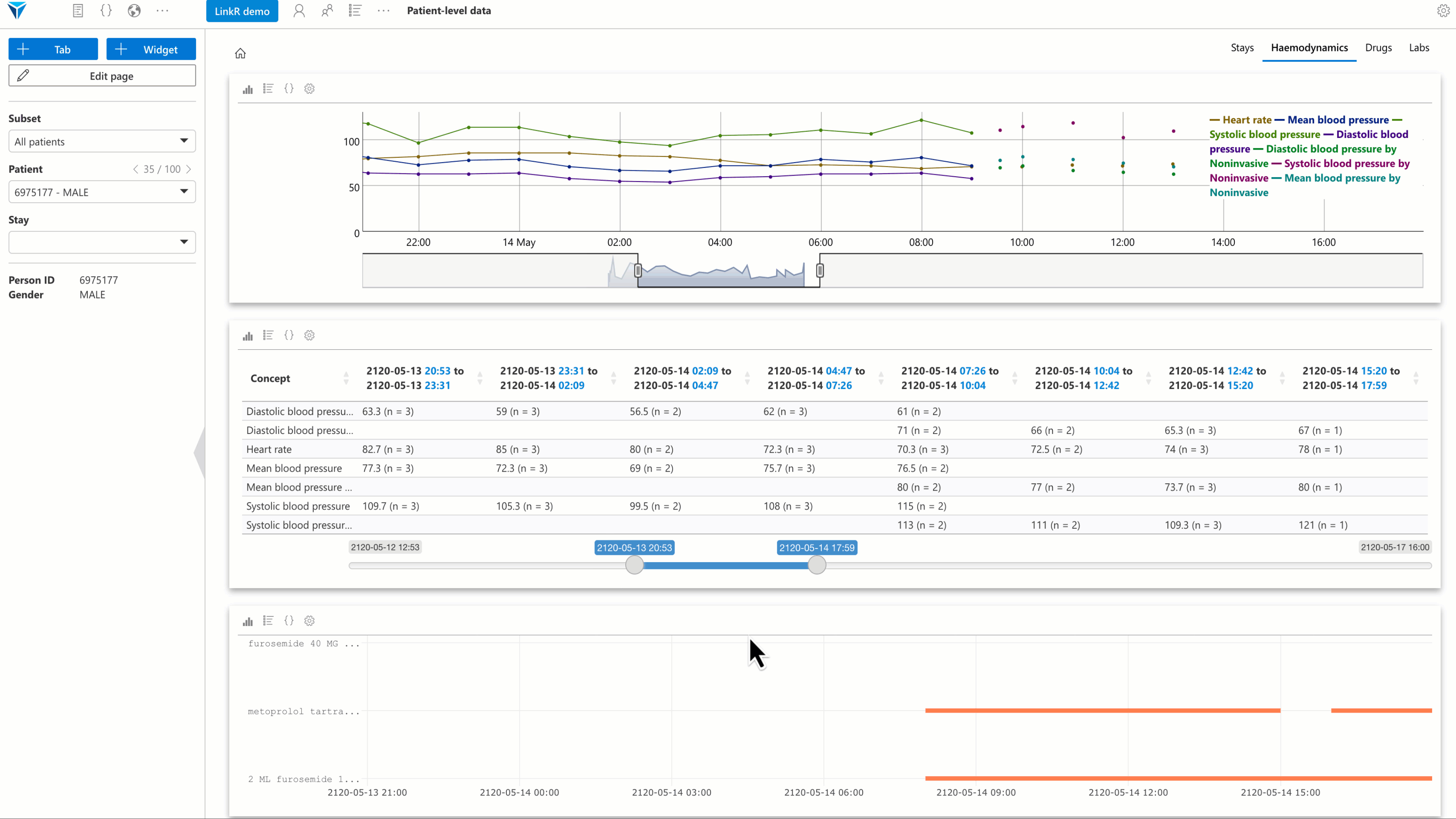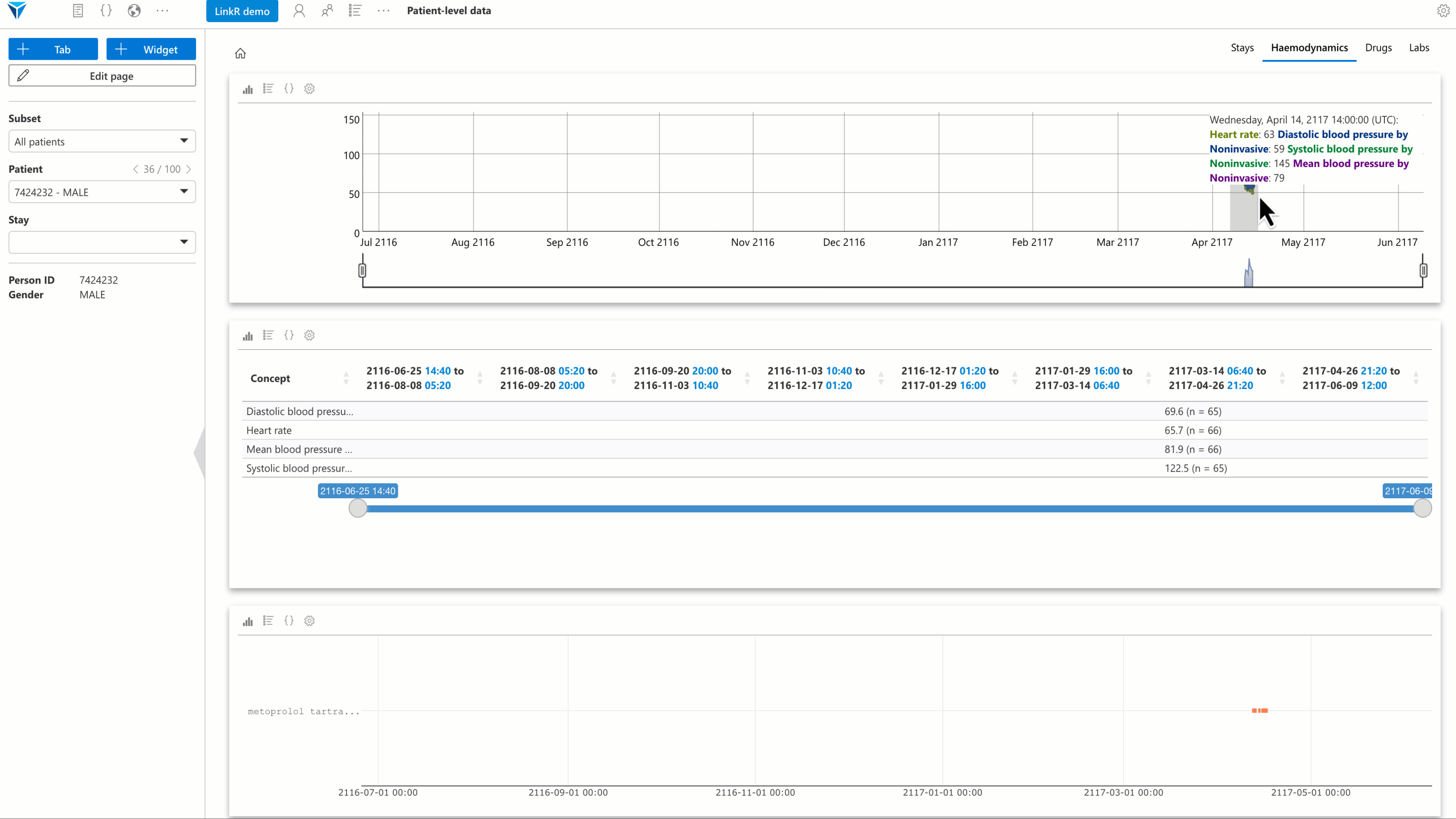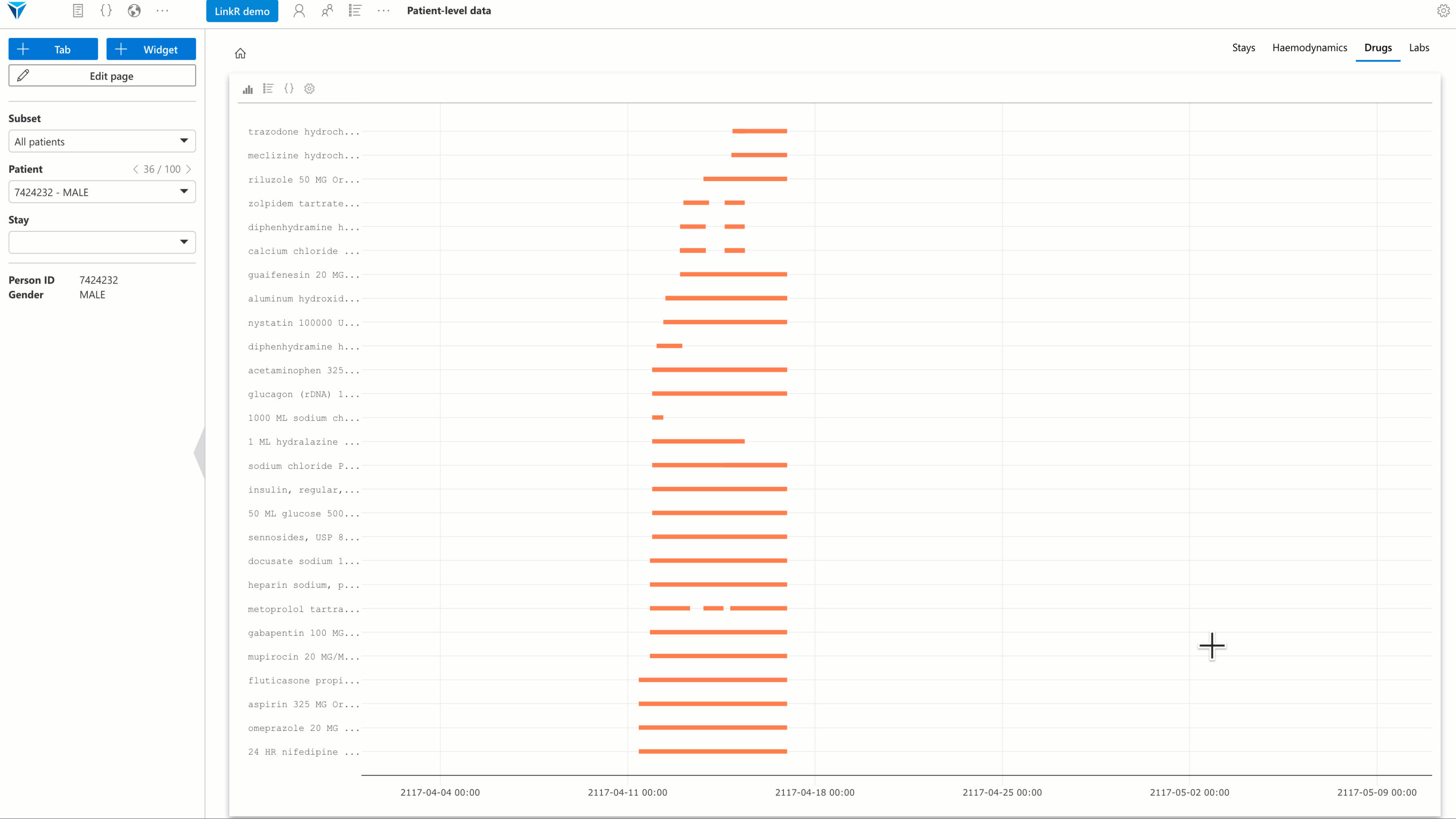
Several plugins have been developed, with a special focus on individual-level data analysis. The goal is to allow users to reconstruct a synthetic and interactive patient record view directly within the research environment.
Examples of available plugins:
- Timeline plugin: visualize data (vital signs, lab results, etc.) as an interactive timeline.
- Text reader plugin: display medical reports and perform full-text search.
- Treatments plugin: view treatments administered to a patient over time.
As always in LinkR, each plugin includes:
- a graphical interface for easy data visualization and filtering
- a programming interface for customization and advanced use
A Template plugin is also available to guide developers in creating new plugins.
Use of DuckDB
LinkR supports importing data from relational databases as well as from CSV or Parquet files.
To ensure maximum interoperability, we chose to rely entirely on SQL queries as the common analysis language.
In this context, DuckDB proved to be an ideal solution: it allows the creation of databases from CSV or Parquet files in seconds, while offering excellent performance. With DuckDB, LinkR can handle tables with millions of rows, even on limited infrastructure.
No matter the data source, it can be explored and analyzed using SQL directly from the platform interface.
Planning for a v2
LinkR is currently written in R, primarily using the Shiny library. While this library allows for fast prototyping of web applications, the developer community around Shiny in R is quite small, which limits contributions to the project.
As an open-source project, we rely on community contributions to keep the project alive.
This leads us to consider a redesign of LinkR as a version 2 using robust and widely adopted web development languages and libraries.
Funding search
InterHop is actively seeking funding to begin development of v2.
We plan to apply for a grant through NLnet.
If you’d like to support us, you can also donate to the association.
Conclusion
One year after Data For Good, LinkR has come a long way: redesigned interface, new plugins, Python integration, enhanced documentation, and a scientific publication.
This project would not exist without the commitment of the many contributors who made it a truly accessible and open-source tool.
With version 2 on the horizon, the adventure continues—and all helping hands are welcome to join in!
InterHop Datathon 2024
The InterHop association is organizing a datathon in September 2024 dedicated to health data, providing participants with access to the MIMIC database in OMOP format. This event offers a unique opportunity to collaborate on data science, data engineering, and artificial intelligence projects applied to healthcare.
Practical information
Participants are encouraged to create their PhysioNet account in advance, including the signature of the Data Use Agreement (DUA).
A kickoff meeting is scheduled for Thursday, August 8 at 1:00 PM, to form teams and refine projects.
You can find all practical information about the datathon in this dedicated article.
Proposed themes and projects
Over the course of 48 hours, several key topics will be explored, ranging from data quality to mortality prediction and health indicator visualization.
1. FINESS+
Update the geographic data of healthcare facilities to ensure interoperability with open systems.
- Main themes: Data engineering, Open data
- Objective: Update and structure the geographic data of the 100,000 healthcare facilities listed in FINESS, making them interoperable with open systems like OpenStreetMap.
- Relevance: Enables cross-referencing of existing databases and improves data quality, with a potential contribution to Toobib.org.
2. Maternity activity indicators
Develop interactive dashboards to analyze and monitor maternity activity indicators.
- Main themes: Data visualization, Data science
- Objective: Create dynamic dashboards to visualize annual maternity activity indicators (C-sections, transfers, epidurals, etc.).
- Methodology: Use of LinkR, an open-source platform that facilitates health data analysis.
- Expected impact: Helps improve maternity unit management and monitor obstetric practices.
3. Data quality
Detect and correct biases in health data
- Main themes: Data cleaning, Pre-processing
- Objective: Produce data quality indicators to assess potential biases from data entry software and improve the reliability of clinical data.
- Methodology: Compare variables from the MIMIC-OMOP database using statistical methods and anomaly detection models.
- Expected impact: Improved health data quality and easier reuse for research.
4. Mortality prediction
Build mortality prediction models to improve patient comparability
- Main themes: Machine learning, Statistics
- Objective: Develop and compare predictive models of ICU mortality (logistic regression, Random Forest, XGBoost, neural networks).
- Methodology: Train models on MIMIC-OMOP and compare them with traditional scores like SOFA and SAPS II.
- Expected impact: Development of an interoperable model applicable to local data, enhancing patient comparability in studies.
5. ICD-10 coding assistance
Assist ICD-10 coding using AI
- Main themes: NLP, Large Language Models
- Objective: Automate the identification of ICD-10 codes.
- Methodology: Use of an approach based on LLMs and RAG (Retrieval-Augmented Generation).
- Expected impact: Improved accuracy and efficiency of medical coding.
Conclusion
This datathon is part of an open science initiative, with the goal of producing interoperable and reusable code for the entire medical and scientific community.
Each project aims to improve the quality, visualization, and use of health data, while encouraging collaboration between clinicians, data scientists, and developers.
The produced source code will be made available on Framagit, and the results may be integrated into platforms like LinkR to be reused and enhanced after the event.